Frequently Asked Questions
GNPS Account¶
I've forgotten my password, what should I do?
If you remember your username, please send an email to ccms-web@cs.ucsd.edu and state your username and request a password reset.
Data Privacy¶
Is the data I upload to analyze publicly visible?
Data you upload to analyze at GNPS will remain private unless you explicitly make it public. The two ways to make your data public are to create MassIVE datasets that make entire datasets public or add individual annotated MS/MS spectra to GNPS spectral libraries.
Will MassIVE datasets that I create be public by default?
No, when creating a MassIVE dataset, it is not public by default. On the input form page, you are able to set a password to access the MassIVE dataset. There is a Make Public link to make your dataset open to the public and documentation can be found here here. Such a feature is very useful because users can create a MassIVE dataset prior to publication submission that is private and password protected. The MassIVE dataset link can be provided to reviewers along with the password so to provide better transparency and increase probability of manuscript acceptance.
Will my GNPS Analysis Tasks be public?
No, by default, when you run an analysis at GNPS (e.g. Molecular Networking) that task will be private only to you. Each task is identified by this 32 task identifier (you can find it in the URL). If you share the URL with a collaborator, they will be able to visualize the results and explore the analysis. We recommend putting these URLs for task analyses in the methods section of your manuscripts.
Data Organization¶
How does the MassIVE data differ from the GNPS spectral libraries?
MassIVE datasets consist of raw data. GNPS spectral libraries consist of spectra that have been individually assigned an identification.
Spectral Libraries¶
How do I get access to the public spectral libraries?
They will be shared with you by default and selected automatically when performing an analysis that requires spectral libraries. This is all included in shared user speclibs. If it does not show up, then do the following.
On the select input files popup, click on the shared files tab, and enter speclibs as an import share.
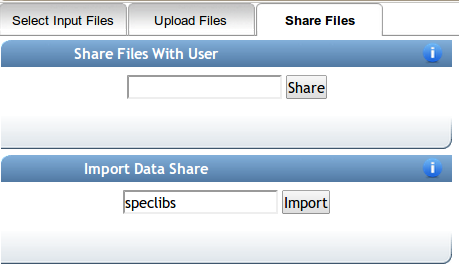
Can I create my own private spectral libraries?
Yes, but it is not recommended and we create these private libraries as a path to making them public. Please contact administrators for further details.
Analyzing Data¶
Can I select more than one file at a time for each group in molecular networking?
Yes. Within each group you can select multiple files, even folders of spectra. Simply highlight the multiple files/folders you want to select and then add to the appropriate group.
What are the advantages and disadvantages of FBMN vs Classical Molecular Networking
Feature Based Molecular Networking (FBMN) provides the following advantages: Area under the curve quantification, Isomer resolution, Filtration of MS/MS that come from chromatographic features, Potential inclusion of other types of edges (e.g. IIN) However, the downsides include: Requires feature finding pre-processing (e.g. MZMine2), Less sensitive to consider all MS/MS than classical, More parameters to set
My Classical Molecular Networking Keeps Failing with the Error - Forcibly terminated by the computational backend
This is most likely the workflow uses too much memory and you have too many spectra in your analysis. Simply increase the minimum cluster size from the default 2, to either 3 or 4. This will be less sensitive but it'll make possible for you analyze your data as a first pass.
Mass Spectrometry Types¶
Does GNPS support data independent acquisition (DIA) data types such as MSe and SWATH?
GNPS natively does not support these types of data with large isolation windows. We recommend that these data be preprocessed so that individual molecule fragmentation can be deconvoluted with tool such as DIA-Umpire or Progenesis QI (MSe) and you can find our documentation here.
GNPS Account¶
How can I reset my password
For now please go ahead and contact our admins via this page.
GNPS Lexicon¶
What is the definition of that word in the context of GNPS ?
Please refer to the GNPS lexicon page. To suggest additional definition, please open up an issue on our GitHub.
Browser Support¶
What Browsers Does GNPS Support
We officially test on the latest Chrome browser. Other browsers, e.g. Firefox, Internet Explorer, Opera, Edge, are not officially supported but likely will not have any issues with GNPS.
Page Contributions¶