FBMN with Progenesis QI
Introduction¶
The main documentation for Feature-Based Molecular Networking can be accessed here. See our article.
Below we describe how to use Progenesis QI with the FBMN workflow on GNPS.
Using Progenesis QI and the Feature-Based Molecular Networking on GNPS¶
Progenesis QI is a proprietary LC-MS feature detection and alignment software developed by Nonlinear Dynamics that is compatible with Waters file format and other proprietary and open mass spectrometry format.
Progenesis QI can perform feature detection, alignment and annotation of non-targeted LC-MS/MS data acquired either in data-dependent analysis (DDA) or MSE data independent analysis (DIA), and can also uses the ion mobility spectrometry (IMS) dimension. Feature-based molecular networking (FBMN) can be performed on any of these data types processed with Progenesis QI.
Citations and development¶
Recommended Citations
This work builds on the efforts and tools from our many colleagues, please cite their work:
Nothias, L., Petras, D., Schmid, R. et al. Feature-based molecular networking in the GNPS analysis environment. Nat. Methods 17, 905–908 (2020).
Wang, M. et al. Sharing and community curation of mass spectrometry data with Global Natural Products Social Molecular Networking. Nat. Biotechnol. 34, 828–837 (2016).
Running Progenesis QI for molecular networking on GNPS¶
For more information on Progenesis QI, please refer to the official documentation at: http://www.nonlinear.com/progenesis/qi/.
1. Import and process the mass spectrometry data¶
- In Progenesis QI (ver. 2.4, Nonlinear Dynamics), import RAW data from Waters QTof data-independent acquisition (DIA) modes such as SONAR, MSE or HDMSE.
- Processed data with Progenesis QI and export the results for GNPS analysis as indicated below. See the Progenesis QI LC-MS tutorial and the tutorial videos for more informations.
2. Export the processing results¶
In the menu, under the “Identify Compounds” tab:
- Select “Export Compound Measurement” to export the feature quantification table (CSV file) containing compound intensity and annotation can be exported (see below).
- Select “Export fragment database” to the export the MS/MS spectral summary (MSP file) containing the list of representative MS/MS spectra (see below). NOTE: Do not select tags to export
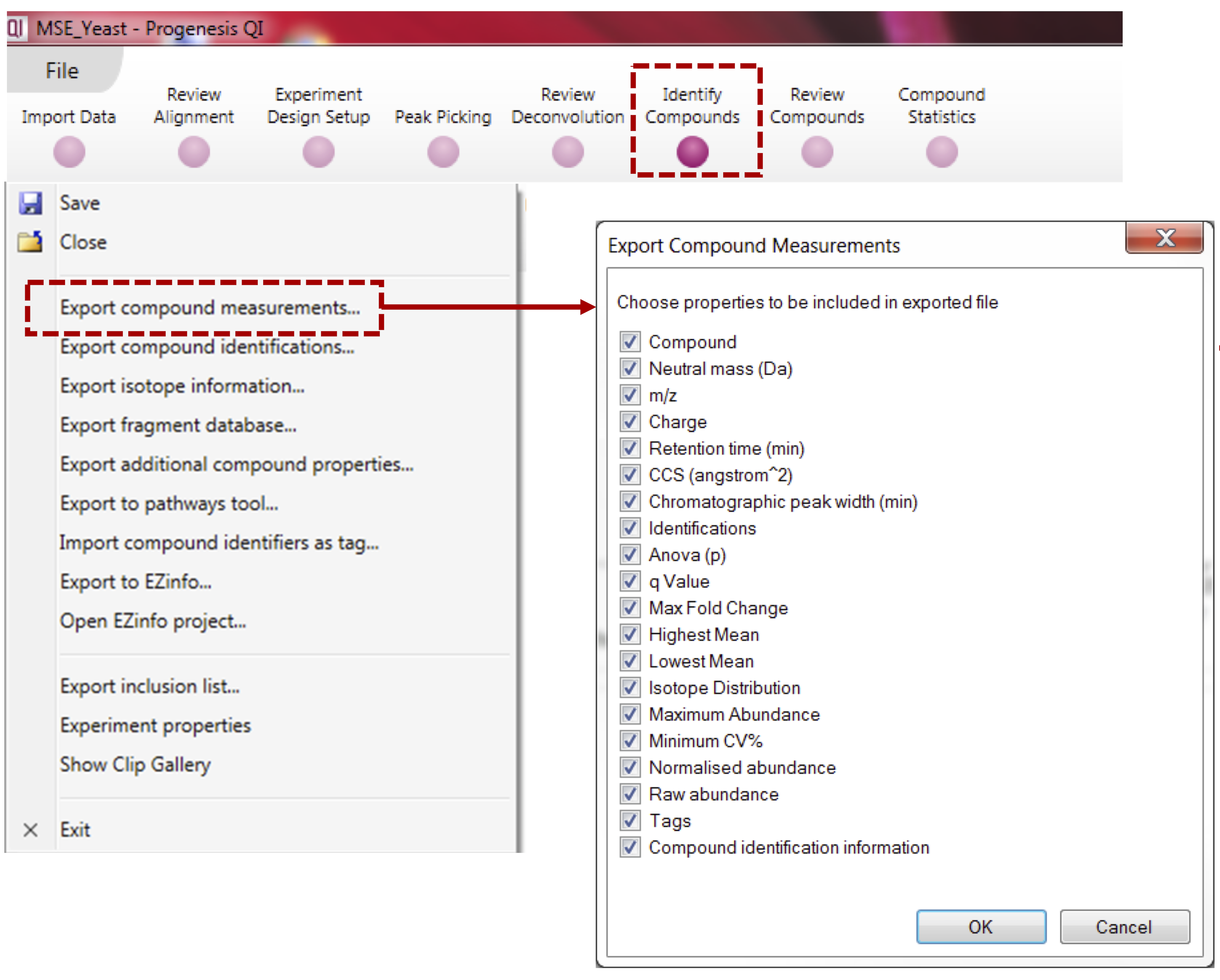
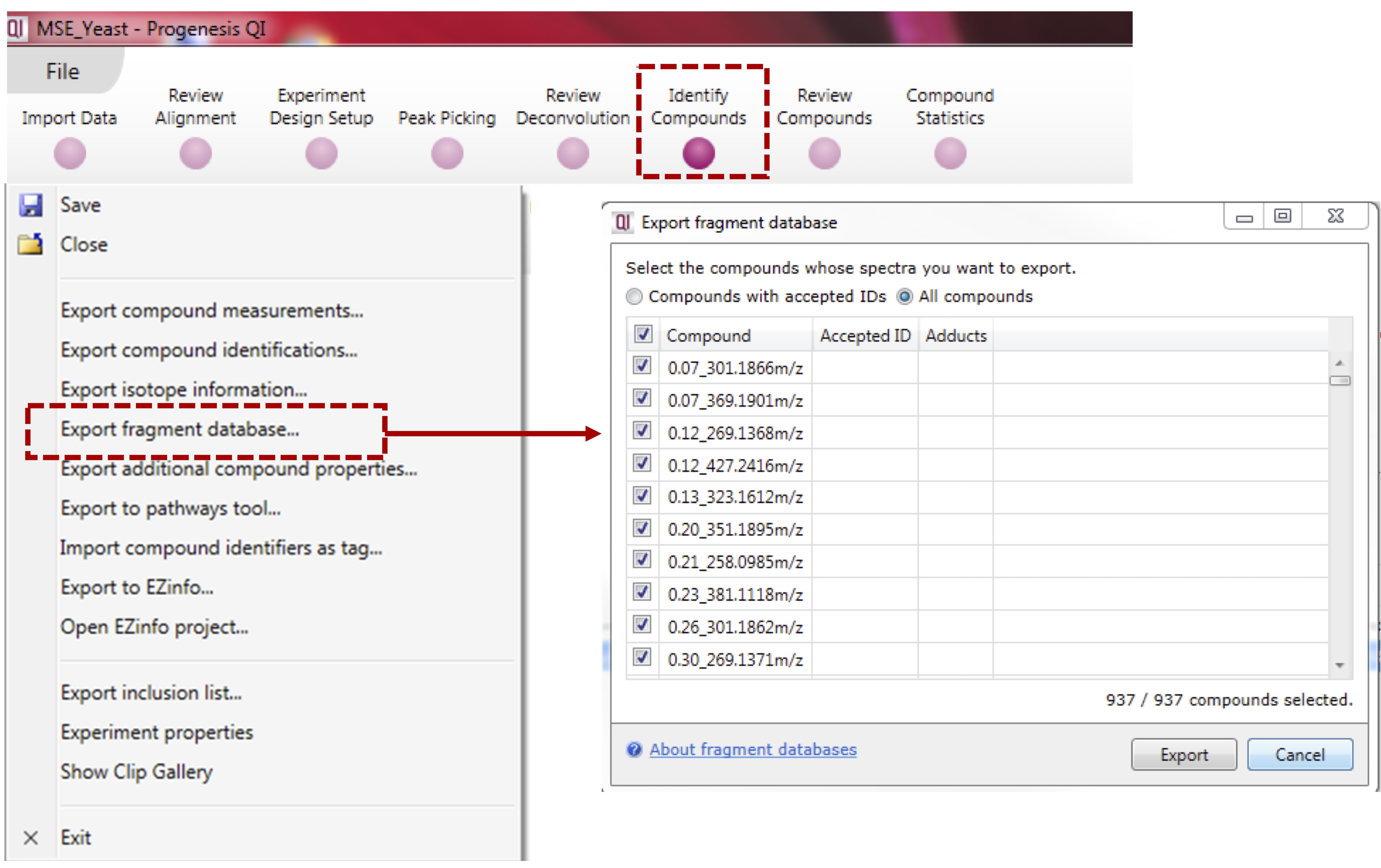
Note
Alternatively, the Symphony software can be used the ion peaks in the entire RAW file can be detected, centroided, and deconvoluted, and converted to a suitable open-source data formats such as Mascot generic format (mgf), mzML, mzXML. More info on in the following note: See more information in the Nov. 2018 Waters Technology Brief. These files can be processed by other softwares supported FBMN, or can be used for classical molecular networking analysis.
Running a molecular network on GNPS¶
FBMN with Progenesis QI results can be performed using the using Superquick or the standard interface:
- Use the Superquick FBMN interface:
- Go to https://gnps-quickstart.ucsd.edu/featurebasednetworking.
- Select the "Progenesis QI" option for the "Feature Quantification Table Source".
- Drag and drop the feature quantification table (.CSV file) into the left box.
- Drag and drop the MS/MS spectral summary (.MSP file) into the right box.
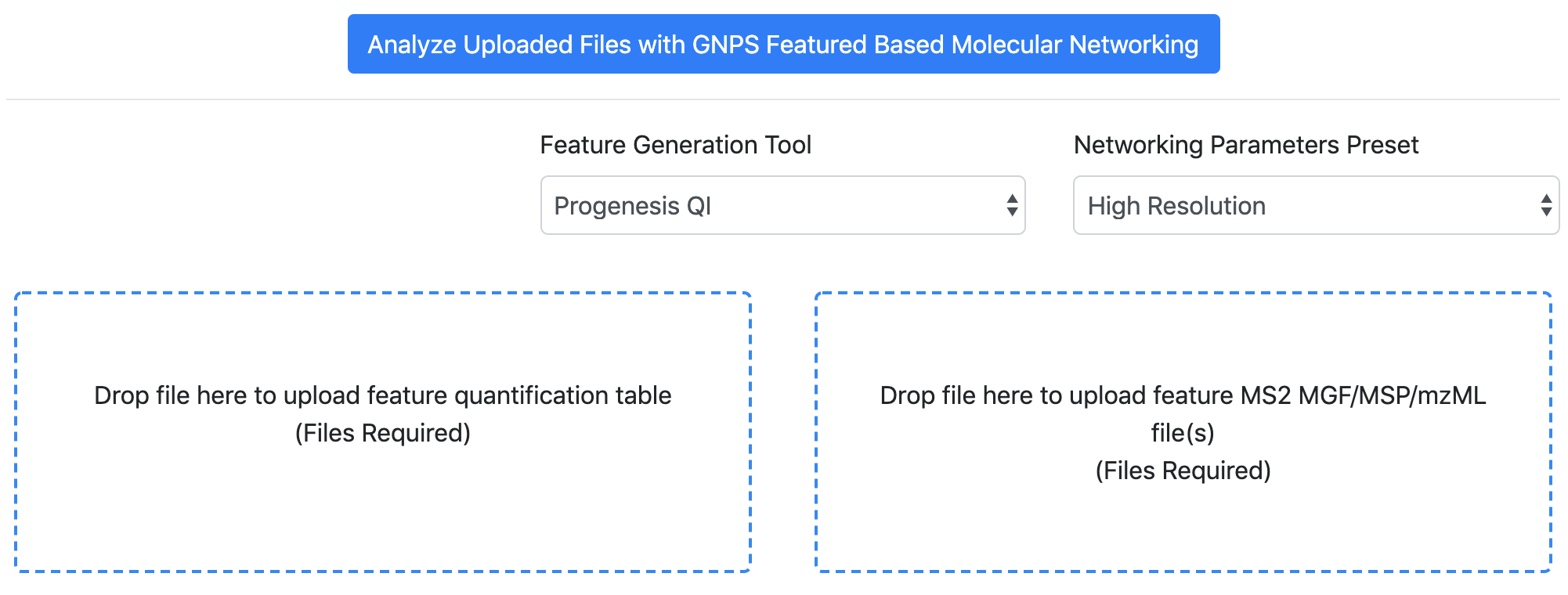
- Use the standard FBMN interface:
- Refer to the main FBMN documentation for more informations.
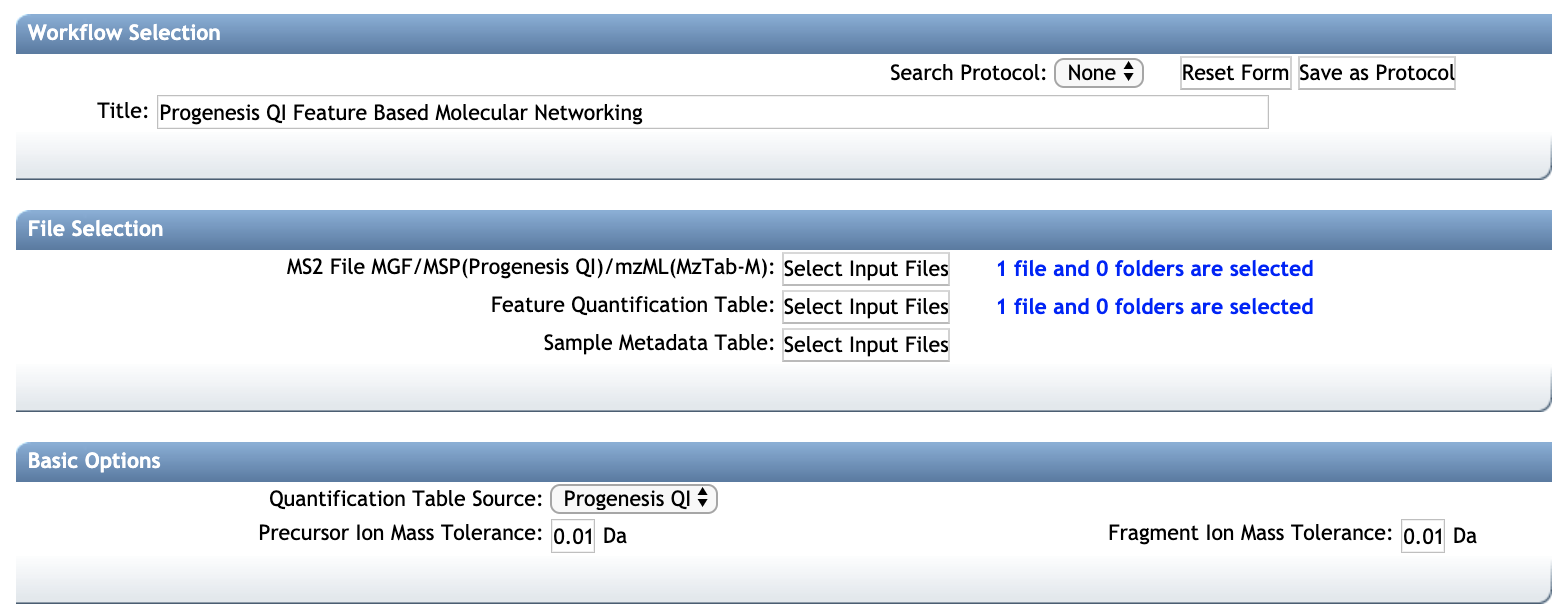
Mapping Progenesis QI annotations in Cytoscape¶
Progenesis QI annotations or metadata can be mapped in Cytoscape as node attribute into the molecular networks. See the Cytoscape documentation for FBMN. For metadata format requirement, see this page.
Helped is wanted to describe in detail the import procedure in Cytoscape.
Acknowledgements¶
Bindesh Shrestha (Waters), Giorgis Isaac (Waters) and Jonathan McSayles (Waters).
Join the GNPS Community !¶
- For feature request, or to report bugs, please open an "Issue" on the CCMS-UCSD/GNPS_Workflows GitHub repository.
- To contribute to the GNPS documentation, please use GitHub by forking the CCMS-UCSD/GNPSDocumentation repository, and make a "Pull Request" with the changes.
Page Contributors¶