Mass Spectrometry File Conversion
Supported Formats at GNPS
GNPS supports mzXML, mzML, and mgf formats for analysis. Our tools do NOT support mzData, xml, raw, RAW, wiff, scan, d, and cdf formats. If some of your files are in these formats, please use the following guide to convert them to open formats supported by GNPS.
There is a specific guide for Waters data conversion. See here
Data Conversion (Super Easy)¶
Drag and drop files on our server here and hit convert.
You will receive a download of all the converted files in a zip file that is ready to be analyzed!
Data Conversion (Easy)¶
This is a complete package for Windows users to convert their vendor formats to GNPS compatible format (mzML or legacy mzXML). It is as easy as putting files into a folder and batch converting them all without any installation (well nearly so).
- Download the zip file here
- Unzip contents onto a folder on your computer (e.g. Desktop)
- Install windows libraries in "pwizLibraries-and-Installation" - Run appropriate program for 32-bit (32-Bit_Double-Click_To_Install.bat) or 64-bit system (64-Bit_Double-Click_To_Install.bat). To find out which type of OS you have please check here
- Put vendor formats in "Input_Files" not embedded in other folders
- Double click on Double-Click_To-Convert.bat - Download zip includes demo files for major vendor formats as a test.
- Wait patiently
- Check all the converted files in Output_Folder
- If there are errors, please check log.txt or read on how to convert files in a more traditional manner.
Data Conversion (Traditional)¶
The .mzXML or .mzML formats are strongly preferred and will be discussed below.
| Vendor | Instrument Software | File Format | Recommended Converter | Notes |
|---|---|---|---|---|
| AB Sciex | Analyst | .wiff | MSConvert | verified |
| Agilent | MassHunter | .d | MSConvert | verified (with issues with scan number export) |
| Bruker | DataAnalysis/Compass | .d | CompassXport | This conversion is through the DataAnalysis software |
| ThermoFisher | Xcalibur | .raw/.RAW | MSConvert | verified |
| Waters | MassLynx | .raw | MSConvert is for full scan/DDA datasets. Symphony is for other modes such as MSe/SONAR/HDMSe/HD-DDA | See more informations about the procedure in this Water Technology Brief |
| Shimadzu | .lcd | MSConvert |
For problems with MSConvert, please contact the ProteoWizard developers.
If you are a vendor and think there are better methods to convert your files to work with GNPS, let us know by contacting us.
Conversion with MSConvert¶
Download and install ProteoWizard from here.
Info
Make sure to choose a Windows-compatible version (includes vendor reader support). You must also have .NET Framework 3.5 SP1 and 4.0 installed.
After installation, from the Start Menu, click the ProteoWizard folder and open MSConvert.
-
Click Browse and select file(s) for conversion. Then click Add to add them to the MSConvert workflow.
-
Choose an Output Directory
-
Under Options, choose mzML (prefered) or mzXML for output format, 32-bit for binary encoding precision and uncheck Use zlib compression.
-
Under filters, choose Peak Picking with Vendor checked, in order to centroid the data. Indicate MS-Levels 1-2. Click Add to add the filter.
Danger
Make sure the PeakPicking filter is the first filter in the list (top position), otherwise the conversion will not be centroided!
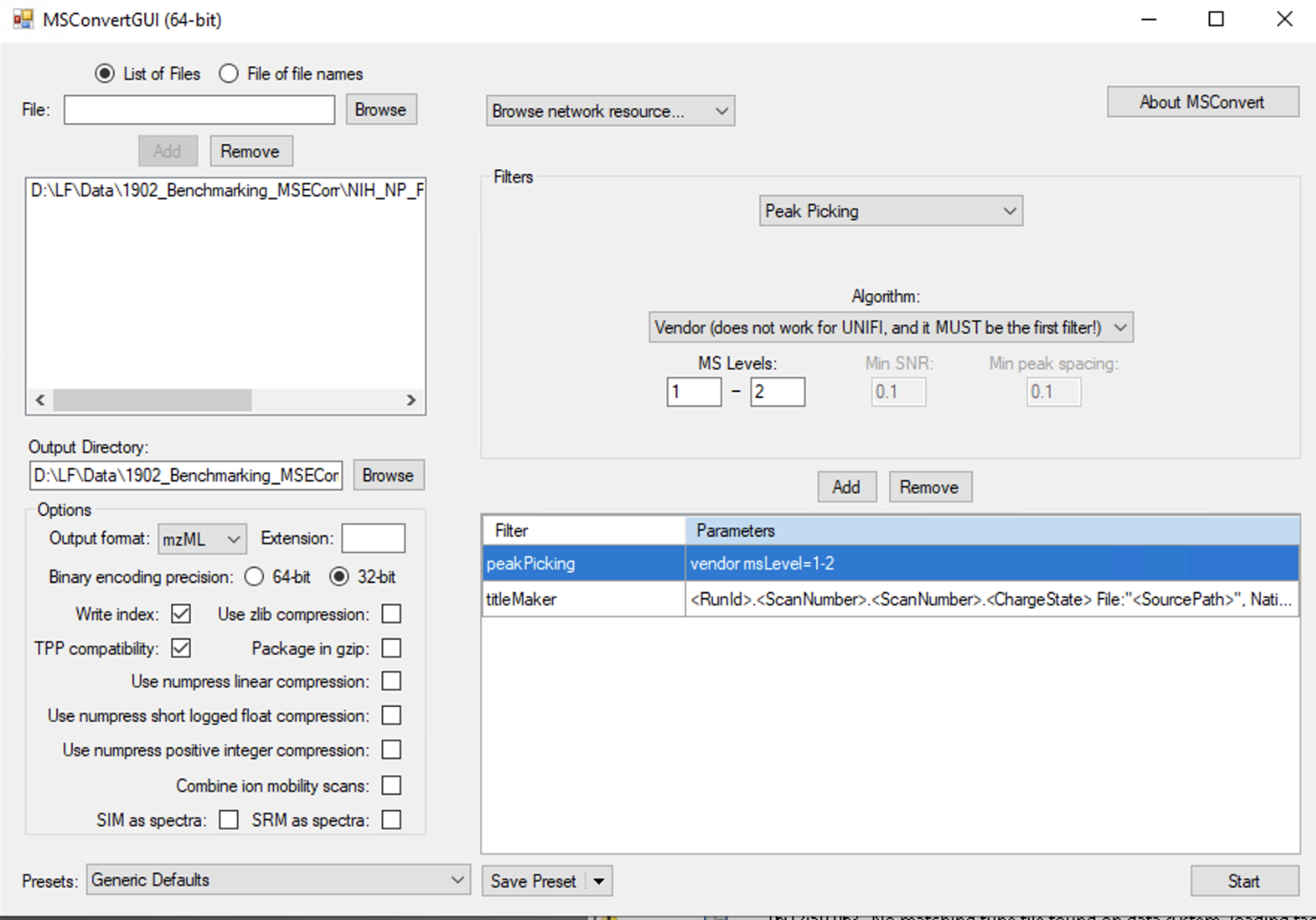
-
Save the parameters for the next conversion. This will save you some time and prevent misconfiguration. In Presets (left bottom), click on Save Presets, and select "Save as default for the format".
-
Click on Start. Check your folder for the new .mzML/.mzXML files. Verify that these files open properly in Insilicos or TOPP View (OpenMS).
Advanced Online Conversion with Proteowizard (MSConvert)¶
It is possible to convert spectrum files online at the sister site - MassIVE. This site is able to handle thermo RAW and Sciex Wiff files. Please check it out here.
Page Contributors¶